Shotgun Metagenomics
Welcome to the realm of shotgun metagenomics, where the invisible becomes visible. At 1010Genome, we're your trusted partner in exploring the genetic diversity of microbial communities through cutting-edge shotgun metagenomics sequencing analysis. Our expertise in this field, paired with state-of-the-art bioinformatics tools, enables you to unveil the secrets within complex ecosystems.
What is Shotgun Metagenomics Sequencing Analysis?
Shotgun metagenomics sequencing is a powerful tool that allows us to uncover the genetic makeup of entire microbial communities in a given environment. It involves sequencing all the genetic material present in a sample, creating a complex mixture of DNA sequences. NGS provides deep sequencing coverage that enables to detect low abundance members of the microbial community that are missed by other conventional methods. Metagenomic provides an opportunity to simultaneously explore two aspects of a microbial community: who is there and what are they capable of doing?

Metagenomics Data Analysis Complexity & Steps

Step-1: Marker Gene Analysis
Marker gene analysis is one of the most straightforward and computationally efficient ways of quantifying a metagenome’s taxonomic diversity. This procedure involves comparing metagenomic reads to a database of taxonomically informative gene families (i.e., marker genes), identifying those reads that are marker gene homologs, and using sequence or phylogenetic similarity to the marker gene database sequences to taxonomically annotate each metagenomic homolog. The most frequently used marker genes include rRNA genes or protein coding genes that tend to be single copy and common to microbial genomes.
Step-2: Assembly
Assembly merges collinear metagenomic reads from the same genome into a single contiguous sequence (i.e., contig) and is useful for generating longer sequences, which can simplify bioinformatic analysis relative to unassembled short metagenomic reads. In some instances, complete or nearly complete genomes can be assembled, which provides insight into the genomic composition of uncultured organisms found in a community

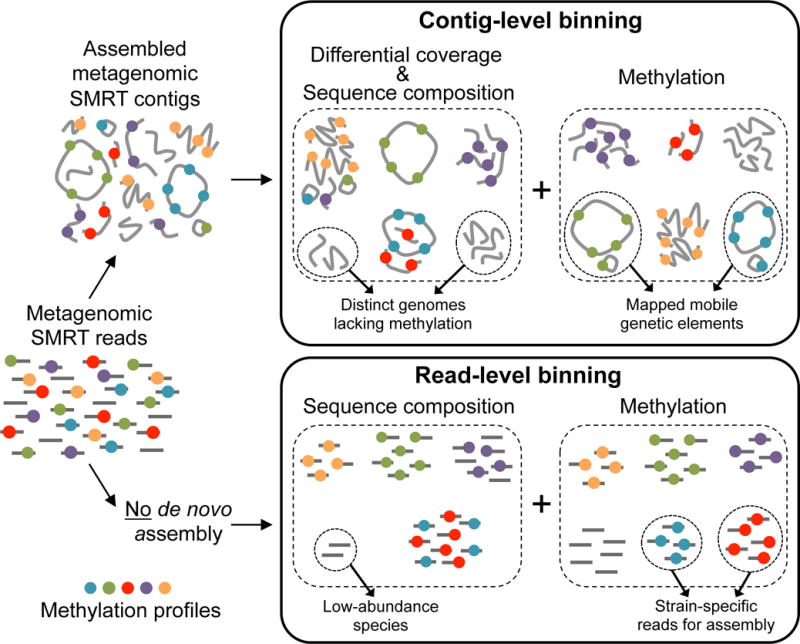
Step-3: Binning
In this crucial step, we group sequences into "bins" representing individual genomes, allowing us to dissect the functional capabilities of specific organisms within the community.
Step-4: Comparative Analysis
We provide insights by comparing your metagenomic data to reference databases or previously sequenced samples, helping you draw meaningful conclusions.

Major Advantages Of Shortgun Metagenomic Sequencing
Shotgun metagenomic sequencing offers several advantages for studying microbial communities:
- Comprehensive Insights: It provides a comprehensive view of all microorganisms present in a sample, regardless of whether they are culturable or well-known, enabling a more holistic understanding of the microbiome.
- Genetic Diversity: It uncovers the genetic diversity within microbial communities, revealing rare and novel species and genes that may be missed by targeted sequencing approaches.
- Functional Potential: Shotgun metagenomics assesses the functional potential of the microbiome, shedding light on metabolic pathways and functional genes present, which is crucial for understanding their roles in ecosystems or host health.
- Hypothesis-Free: Unlike targeted approaches, shotgun metagenomics is hypothesis-free and does not rely on prior knowledge of the microbial community, making it suitable for exploratory research.
- Comparative Studies: It enables comparisons between different samples, facilitating the study of microbiome variations in response to environmental changes, diseases, or other factors.
- Discovery of Novel Microbes: It often leads to the discovery of new microbial species and strains with potential applications in biotechnology, medicine, and more.
- Complex Community Analysis: Shotgun metagenomics can assess complex microbial communities in diverse environments, including the human gut, soil, water, and extreme ecosystems.
- Flexibility: Researchers can apply shotgun metagenomics to a wide range of samples, making it a versatile tool for various scientific disciplines, from ecology to genomics.
Our Shotgun Metagenomics Deliverables
Unleash the power of shotgun metagenomics and embark on a journey of discovery with 1010Genome.
Our shotgun metagenomics sequencing analysis services are your gateway to exploring the hidden diversity and functional potential of microbial communities.

